I first heard of kanban ages ago as a passing mention on a tech podcast I was listening to. If you’re unfamiliar, it is a lean method for streamlining manufacturing that originated at Toyota. It has been widely adopted in the tech industry for software development.
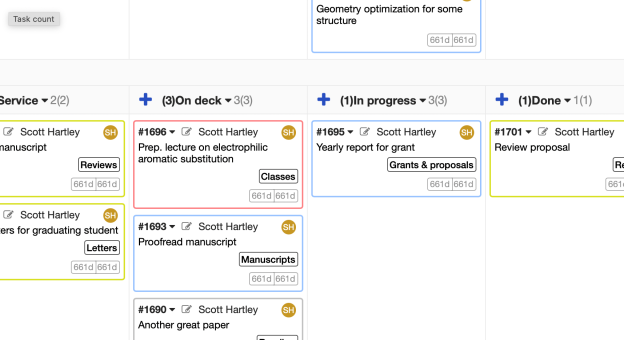
Very briefly, the core of the method is a kanban board, which is probably most easily visualized as a large table with Post-it notes that can be moved around. Each note represents a work item. They are put in columns that indicate their status (e.g., “To do”, “In progress”, “Finished”). Given its history as a way to coordinate teams, I tried to use the kanban method to help manage my research group. But to be honest it really never clicked. It was more of an imposition on my students than something that was genuinely useful.
What has been very useful is using a private kanban board to manage my personal productivity. This method is based on the book Personal Kanban by Jim Benson. I messed around with various todo list methods for years and none of them worked for me. The personal kanban method has been genuinely life changing, however.
Continue reading